
Bridge preclinical findings to clinical outcomes with proteomics
Measure proteins across patient samples. Quantify with consistency. Identify signals that translate to the clinic.
The Nomic Platform enables scalable, quantitative proteomics for translational studies
Absolute Quantification
Translate measurements directly across samples and studies.
Flexible Design
Build protein signatures from broad discovery to focused patient stratification.
Curated, Biology-Driven Targets
Measure biologically relevant targets prioritized for clinical and translational impact.
Proteomics advancing
translational research
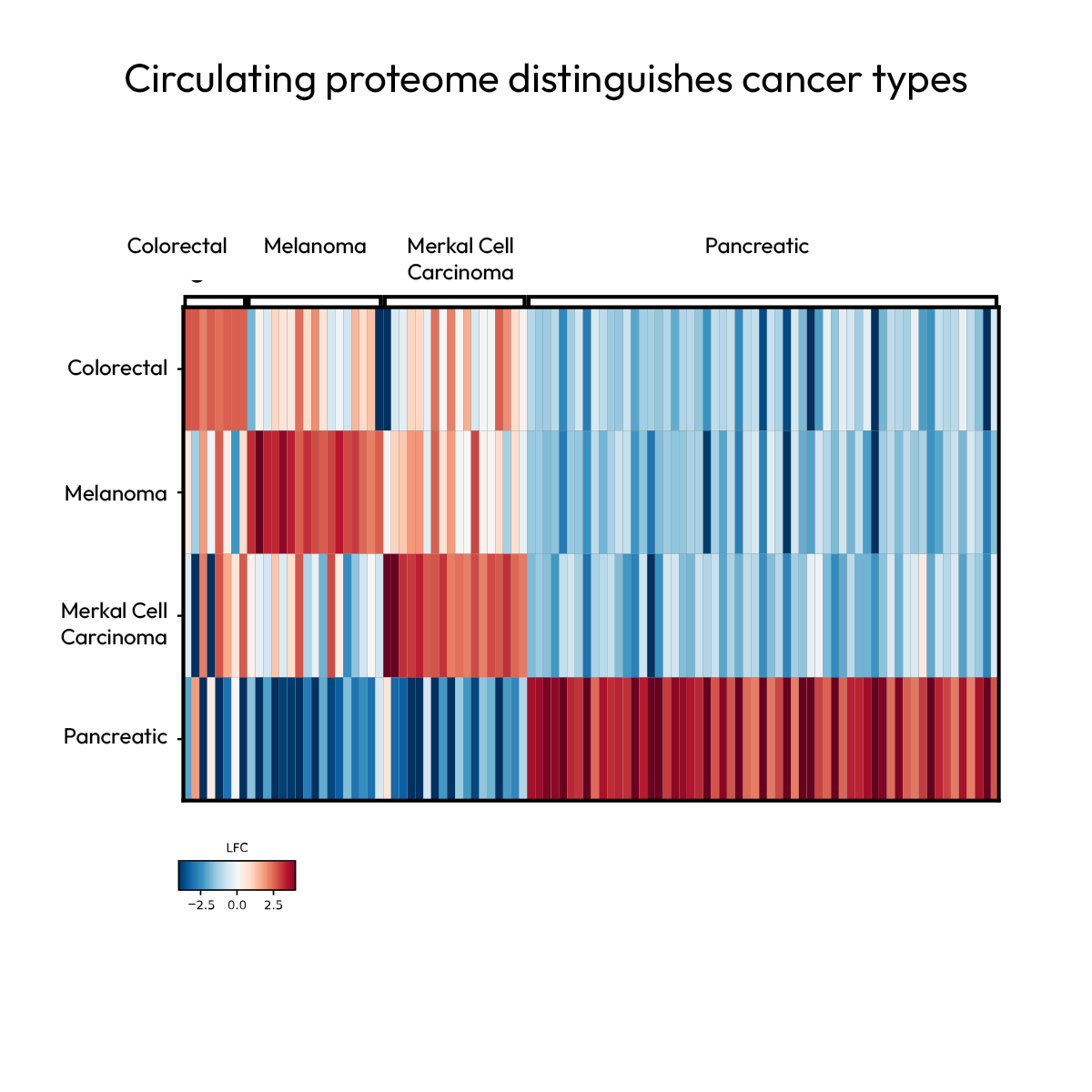
Profiling protein signatures for patient stratification and treatment response
Detailed understanding of patient heterogeneity is crucial for patient stratification. The Nomic Platform effectively captures protein signatures associated with disease and therapy response across patient samples.
Across multiple pan-cancer studies, Nomic’s protein-level data was used to segment patients by cancer type, revealing distinct phenotypes across colorectal, melanoma, Merkel cell carcinoma, and pancreatic cancer, and identified signatures associated with response to checkpoint inhibitor therapy through longitudinal sampling.
Data generated in collaboration with the Bod Lab at Massachusetts General Hospital Cancer Center
Decoupling therapeutic response from adverse events
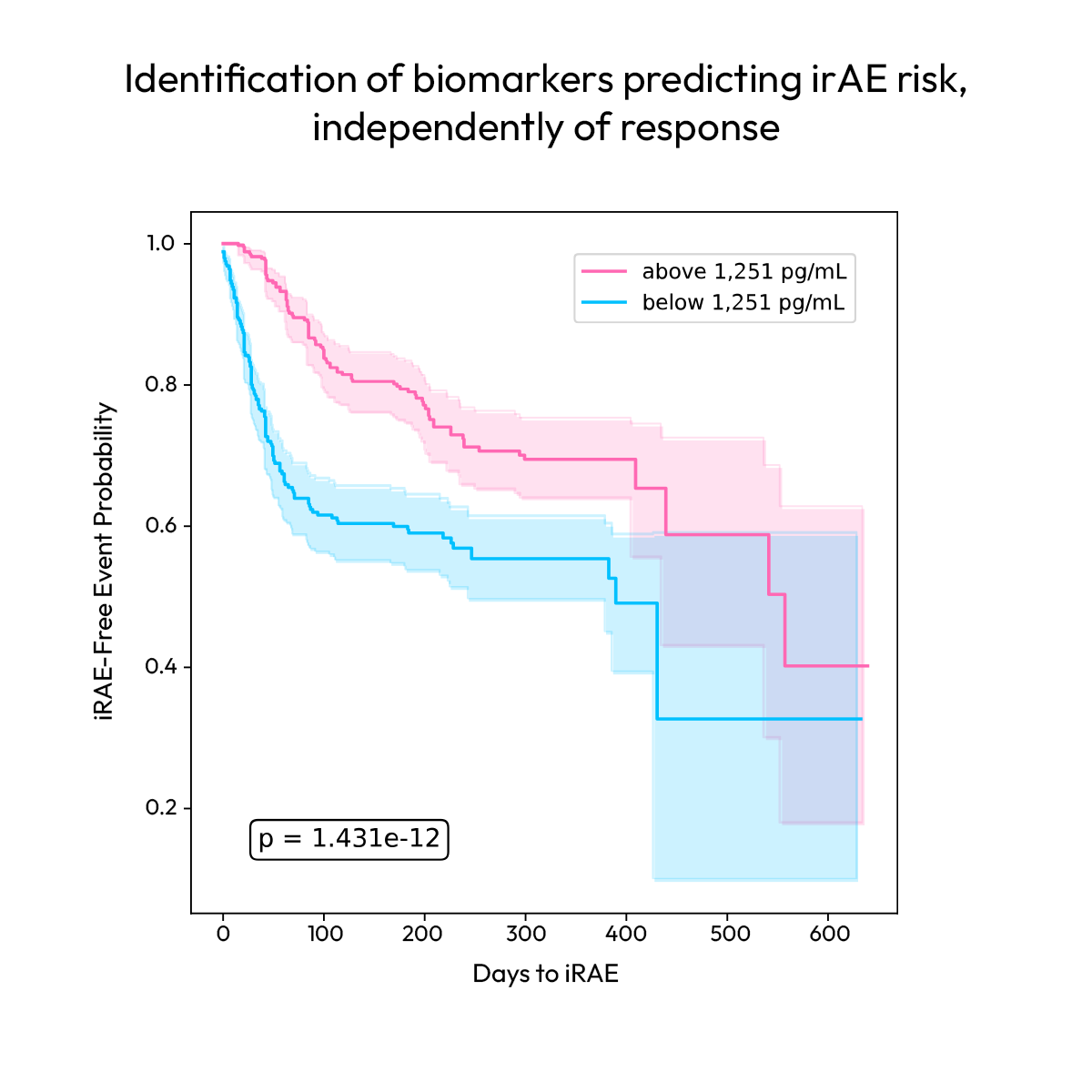
Distinguishing therapeutic response from adverse events requires biomarkers that resolve overlapping biology. In the largest-scale plasma proteomic study to date in patients receiving checkpoint inhibitor therapy, the Nomic Platform captures protein signatures decoupling response from immune related adverse events.
Protein-level data reveals that response and toxicity signatures are largely independent, enabling identification of patients likely to benefit without elevated safety risk, as well as those requiring additional monitoring. Distinct patterns across cancer types and treatment modalities further highlight the need for broad, well-powered profiling to capture clinically relevant biomarkers and support more precise patient stratification.
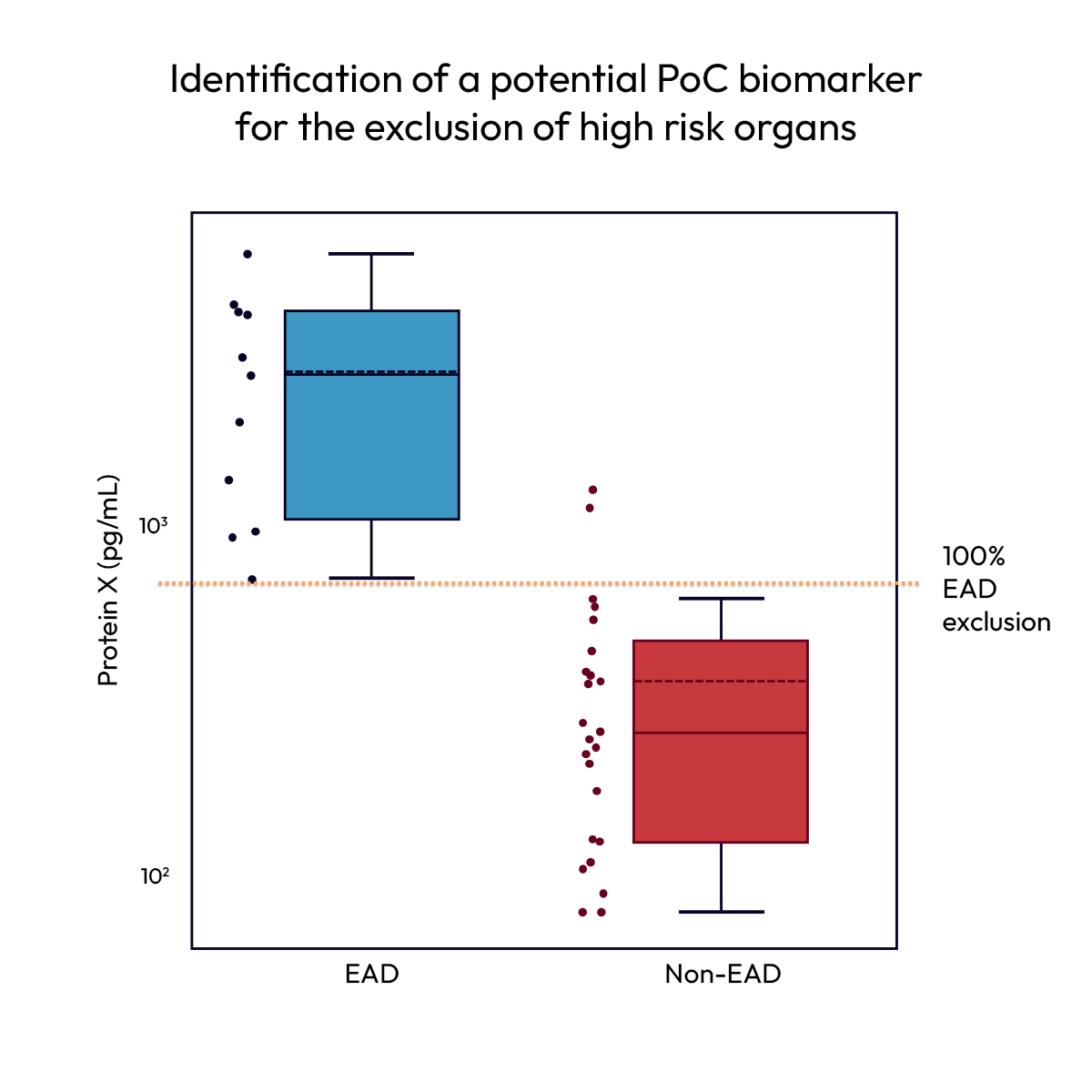
Predicting transplant
outcomes from pre-implantation biomarkers
Identifying transplant viability requires biomarkers that reflect organ health prior to implantation. In proteomic profiling of liver perfusate samples, Nomic’s Omni 1000 effectively identifies protein signatures associated with transplant outcomes.
Protein-level data reveals markers of cell stress and tissue damage enriched in livers that develop early allograft dysfunction. A single biomarker enables accurate exclusion of high-risk organs while retaining the majority of viable grafts, supporting more informed transplant decisions.
Data generated in collaboration with the Tara Sigdel, PhD, and the Sarwal Lab at UCSF.
Proteomics that supports you all the way
From discovery screening to drug R&D and clinical studies, Nomic provides a service that scales with your projects. Whether using Omni 1000, context-specific Core options, or even a tailored Flex panel designed to your objectives, you can adapt your approach without compromising data quality or cross study compatibility.
Discovery
- Target ID
- Therapeutic Biomarker Discovery
- Indication Selection
- Target Validation
- Perturbation Biology
- MoA Studies
- Early Tox Profiles
Drug R&D
- Target Engagement
- Hit-to-Lead
- Hit Optimization
- ADME/Tox
- Pre-Clinical/IND Enabling
Translational
- Biomarker Validation
- Patient Stratification
- PK/PD
- Safety
- Disease Prognosis
Stop inferring biology. Start measuring it.
See how quantitative protein data drives decisions from target selection to patient stratification.


Resources for discovery teams
Your research project deserves a protein answer
Our scientists have designed proteomics studies across target ID, perturbation biology, MoA, and early toxicity. Share the specifics of your program, and we will tell you what is possible.

